 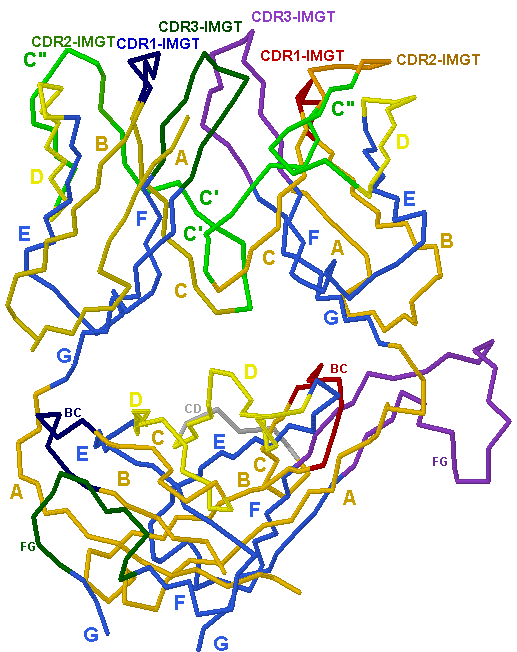
We illustrate the usefulness and application of this methodology by the analysis of the human TR repertoire response towards a model immune challenge, the H1N1 vaccine. This approach consists of 5′ rapid amplification of cDNA ends (RACE) 12, 26 to avoid amplification bias associated with multiplex PCR, 454 sequencing to bypass the limitations of short-read assembly and IMGT/HighV-QUEST analysis 21, 22 to ensure the highest quality in sequence interpretation of full-length rearranged human TR V-BETA transcripts. We now introduce a high throughput methodology for a standardized comparative TR analysis on the basis of IMGT-ONTOLOGY concepts. IMGT/HighV-QUEST uses the same algorithm as IMGT/V-QUEST and achieves the same degree of resolution and high quality results. ImMunoGeneTics (IMGT)/HighV-QUEST 21, 22 ( ) is the authentic high throughput version of the IMGT/V-QUEST tool 23, 24, 25 (acknowledged as the international reference for immunoglobulin and TR sequence analysis, CSH Protocols, WHO/IUIS). Earlier studies also explored various bioinformatic tools but different algorithms added potential layers of discrepancy. 454 sequencing was also used previously 16, 17, 19, 20 but only in combination with multiplex PCR. 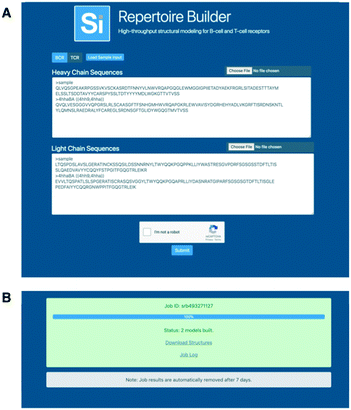
In these studies, Illumina sequencing was primarily used, with a major advantage of generating very deep data, but a disadvantage that the read length was short and the data either required assembly 12, 13 or focused exclusively on the CDR3 (refs 14, 15). The complexity and depth of the human TR repertoire was recently explored in several studies using next generation sequencing (NGS) 12, 13, 14, 15, 16, 17, 18, 19, 20. Classical DNA cloning and Sanger sequencing techniques are laborious and generally limit data to a few hundred, or in rare cases a few thousand, TR sequences per investigation 6, 7, 8, 9. The approach by tetramer staining or antigen-induced cytokine release 6, 7, 8, 9, 10, 11 is limited by our knowledge of mapped epitopes. It remains a significant challenge to understand the diversity and specificity of T cells, particularly during natural infection. The specificity of the TR predominantly depends on the CDR3 created by the V-(D)-J rearrangement. When docking its cognate pMH ligand, the CDR1 and CDR2 facilitate binding of the receptor to the MH helices, while CDR3 principally engages the peptide within the MH groove 4, 5. Each V domain comprises three highly flexible complementarity-determining regions (CDR) at the antigen-binding face of the receptor 1. The C region of the TR chains is encoded by the TRAC and TRBC ( TRBC1 and TRBC2) genes, respectively 1. The V-ALPHA domain results from the rearrangement between a TRAV gene and a joining J ( TRAJ) gene, whereas the V-BETA domain results from the rearrangement of a TRBV gene, a diversity D ( TRBD) gene and a joining J ( TRBJ) gene. Each chain comprises a variable (V) domain and a constant (C) region that includes a C domain and connecting, transmembrane and cytoplasmic regions. The αβ TR is a membrane-bound, clonotypic, heterodimeric protein comprising one alpha chain (TRA) and one beta chain (TRB) 1. The TR repertoire is vast, with direct estimates of 2.5 × 10 7 unique αβ TR per individual 2 and significantly higher numbers by theoretical calculations 3. The T cell receptor (TR) 1 is critical for peptide/major histocompatibility (pMH) recognition. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
